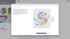
We’re thrilled to announce the added capabilities in our latest ROSALIND update to v3.15, including expanded COVID-19 support and resources, as well as significant enhancements to our Pathway interpretation engine.
What's new in the COVID-19 Community?
The SARS-CoV-2 genome is available for new experiments and to support the work within the COVID-19 Community Space. Much of our focus these past few months has been to further facilitate the analysis and discovery into the viral gene expression of SARS-CoV-2. Many of the recent NCBI public data submissions with SARS-CoV-2 samples and COVID-19 patients have already been processed and added to the Community Space.
ROSALIND Pathways has added a new Virology category and collection of Knowledge Bases to enhance COVID-19 research. New entries include Pathogen-Host Interactome (P-HIPSTer) and GEO Virus Perturbations. Insights from the new Virology Collection can be added to all existing experiments by simply adding a new filter, or comparison.
Learn more about the COVID-19 Community at this link: https://www.onramp.bio/blog/coronavirus
Enhanced Pathway Interpretation
Pathway interpretation has been upgraded now with over 50 Knowledge Bases and up-to-date versions of every database available. New categories intelligently group and organize each Knowledge Base into categories that streamline exploration and focus research, including: Pathways, MSigDB Pathway Collection, Oncology & Immunology, Transcription & Regulation, Diseases, Drugs, Virology, Ontology Collection, Cell Types & Tissues and Protein Collection. Additionally, the Ontology Collection now features Advanced Pruning (Elim) to eliminate largely redundant terms.
- Pathways: WikiPathways, BioPlanet, Reactome, Panther, BIOCYC, Pathway Interaction Database, and the Small Molecule Pathway Database
- MSigDB Pathway Collection: Hallmark, Chemical and Genetic Perturbations, MSigDB-Reactome, MSigDB-BioCarta, MSigDB-Protein Interaction Database
- Oncology & Immunology (MSigDB): Cancer Gene Neighborhoods, Cancer Modules, Oncogenic Signatures, Immunological Signatures
- Transcription & Regulation: MSigDB-Transcription Factor Targets, JASPAR-Transcription Factor Targets, TRRUST-Transcriptional Regulatory Networks, miRNA Targets, miRNA Target Interactions, miRNA Target Predictions, and Chromosome Location
- Diseases: ClinVar, PheWeb, GWAS, DisGeNet, Jensen Diseases
- Drugs: FDA Approved Drugs (DSigDB), Kinase Inhibitors (DSigDB), BROAD Connectivity Map (DSigDB), Computational Drug Signatures (DSigDB), Guide to Pharmacology, Drug-Gene Interaction Database, Drug Matrix - Toxicogenomics Gene Signatures
- Virology: P-Hipster - Pathogen-Host Interactome, GEO - Virus Perturbations (Up & Down)
- Ontology Collection: Human Phenotype Ontology, Biological Processes, Molecular Functions and Cellular, Molecular Functions and Cellular Component, GO Molecular Function (MSigDB), GO Biological Processes (MSigDB), and GO Cellular Component (MSigDB) - Note that Advanced Pruning is now available (p-Elim) to eliminate largely redundant terms in Gene Ontology
- Cell Types & Tissues: Human Cell Atlas
- Protein Collection: Protein-Protein Interactions, Interpro, Pfam, SMART, GENE3D, and Prosite
For new users, you can choose the right plans for your team from our online catalog (https://rosalind.onramp.bio/store/catalog), sign up for a Scientist Trial or learn more about Enterprise Subscription Plans by contacting us here: https://www.onramp.bio/contact-us
Related Posts
-
For all Research affected by Social Distancing: Remote Science in the Era of Social Distancing demonstrates how working remotely does not mean that science must stop.
-
For Scientists on ROSALIND: Collaborating on Genomic Data is a walkthrough on how to set up real-time collaboration on genomic data (like Google Docs). Read the post here
-
For Directors and Managers: The Value is in Collaboration (not the pipeline) discusses how collaboration is key to empowering research in your lab.
-
For every Researcher, Public Data exploration shows how we have made it 100x easier to work with public data than ever before.
- To learn more about our latest capabilities, Empowering Scientists with the ROSALIND Platform is an overview of all analysis types, species and features currently available to the general public on ROSALIND.
Understanding the immediate economic shifts happening around the globe, we are offering special pricing on our subscriptions with coupon code: REMOTESCIENCE for a 33% discount on monthly subscriptions through the rest of 2020.